All published articles of this journal are available on ScienceDirect.
Isolation and Characterization of Phosphate Solubilizing Bacteria from Phosphate Solid Sludge of the Moroccan Phosphate Mines
Abstract
Background:
Although phosphorus (P) reserves in the soil are important, orthophosphate ions are the only forms of P that can be used by plants. The specificity of phosphate solubilizing bacteria (PSB) to convert insoluble forms of P to an available form Pi is an important trait to reactivate immobilized phosphorus. Therefore, the use of PSB as inoculants increases the P uptake by plants.
Objective:
The present study aims at isolating and selecting the phosphate solubilizing bacteria from Phosphate Solid Sludge (PSS) in order to improve the agronomic efficacy of P fertilizers.
Materials and Method:
The strains isolated from 7 samples of phosphate sludge were tested for their ability to solubilize phosphate in NBRIP medium with Tricalcium Phosphate (TCP) as the sole source of phosphorus, strains with a high solubility index (PSI) were selected. The isolates obtained have been studied for their ability to solubilize TCP quantitatively, and their phosphate solubilizing activity was estimated by the vanadate-molybdate method. The production of Indol-1-Acetic Acid (IAA), Siderophore, and Hydrogen Cyanide (HCN) by the isolates was tested. The best strains have been identified based on 16S rRNA gene sequence comparisons.
Results:
27 strains were isolated on selective medium NBRIP from Phosphate solid sludge. They all showed significant different abilities to solubilize P. CB19 was the most efficient strain in NBRIP agar with SI= 4.79; BM11 was the most efficient strain in NBRIP liquid medium releasing 174.33μg/ml soluble P. Based on 16S rRNA gene sequence comparisons, four genera were identified: Pseudomonas, Serratia, Pantoea and Enterobacter.
Conclusion:
The PSB isolates varied in their Phosphate solubilizing efficiency, IAA production, Siderophore, and HCN.
1. INTRODUCTION
Phosphorus (P) is one of the major essential macronutrients for growth and plant development. It is involved in key plant metabolic processes such as storage of cellular energy, photosynthesis, and respiration [1]. The majority of agricultural soils contain a high reserve of P. However, the amount of soluble P in soil is generally very low (0.4-1.2 g/kg) [2, 3]. The important phosphate reserve in the soil is habitually accumulated as a consequence of regular applications of P fertilizers. According to FAO [4], more than 175.5 million tons of chemical fertilizers are used in agriculture to obtain the best yield of crops and therefore cover increasing habitat needs. More than 80% of P fertilizer, applied to the soil, becomes unavailable for plant via sorption or precipitation by reaction with Ca2+ in calcareous soils, Al3+ and Fe3 + in acidic soils [5].
At present, the main objective of soil phosphorus management is to optimize agricultural production and minimize phosphorus losses in soils. Insoluble P compounds can be solubilized by organic acids, phosphatase enzymes and complexing agents produced by soil microorganisms [6]. Hence, among the solutions of soil phosphorus management are the use of Phosphate solubilizing microorganisms as soil inoculums to enhance chemical fertilization and thereby improve plant growth and yield [7, 8]. For several decades, agricultural microbiologists have studied the ability of some bacteria to dissolve insoluble P [9]. Currently, most bacteria named PSB belong to the genera: Pseudomonas, Serratia, Burkholderia, Achromobacter, Agrobacterium, Micrococcus, Aerobacter, Flavobacterium, Acinetobacter, and Pantoea [9-13].
Morocco covers a significant share in global phosphate reserves; it contains 75% of phosphate reserves on the planet, and it is the leading exporter of phosphate and its derivatives. The country’s global market share is more than 30% [14]. Phosphate deposits in Morocco are located in three areas - Khouribga area (Oulad Abdoun Plateau), Gantour area (Youssoufia area) and Laayoune-BouCraa [14]. The phosphate processing activity generates enormous solid sludge. The chemical characterization of PSS has been the subject of several works in the past, and they consist generally of clay, but the phosphate material is found there. The presence of phosphorus in these PSS means a loss of phosphates during treatment. OCP Group aims to develop phosphate as part of a vision of sustainable agricultural development and respect for the environment. Consequently, the search for appropriate solutions to the pollution risk posed by this PSS and the recovery of some of the lost phosphorus becomes necessary.
In this context, the objectives of the present study are to isolate and characterize phosphate-solubilizing strains from Phosphate solid sludge and assess their PGPR traits in order to evaluate the phosphate solid sludge.
2. MATERIALS AND METHODS
2.1. Sample Collection And Analyses
PSS samples were collected in seven different locations (Table 1) in Khouribga region from the rhizosphere of Tamarix sp. (S-BT), Nicotiana glauca (S-BN), Atriplex halimus (S-AT), Medicago Arabica (S-BM), Acacia Cyclops (S-ACY), and one sludge sample was collected far from the vegetation (S-CB and S-AC). The samples were collected at a depth of 10 to 40 cm and they were mixed to obtain a representative sample. The samples were transported to the laboratory and conserved at 4 °C for further analysis.
Physico-chemical characteristics of collected PSS samples such as moisture content, pH, and organic matter, and the enumeration of total Bacteria were also determined (Table. 2). The pH was measured in 1:2.5 soil: water suspension. Organic matter was determined by the dichromate-oxidation method [15]. Total bacteria were enumerated by the plate counts method. 1 g dry weight of samples suspended in 9.0 ml of phosphate buffer saline (pH=7.2) was used to provide a dilution series from 10-1 up to 10-7.
2.2. Selection and Purification of Phosphate Solubilization Bacteria
One gram of each PSS samples was suspended in 9.0 ml of phosphate buffer saline (pH 7.2) and serial dilutions from 10-1 up to 10-6 were realized. 0.1 ml suspension was spread on NBRIP solid medium (10 g/l D-glucose, 5 g/l MgCl2 6H2O, 0.25 g/l MgSO4 7H2O, 0.2 g/l KCl, 0.1 g/l (NH4)2SO4), amended with 5.0 g/L Tricalcium Phosphate (TCP) as a sole source of P [16]. It was incubated at 30°C for 7 days. The bacterial colonies with a clear halo zone were selected and purified three times on NBRIP solid medium. The purified bacterial isolates were maintained on nutrient Agar Plates and stored at 4°C, and a copy of each isolate was stored as a glycerol 40% stock at -30°C.
2.3. Inoculum Preparation
The purified isolates were grown in the NBRIP medium. Each isolate was inoculated into test tubes containing 3 ml of NBRIP liquid medium and incubated for 24 h at 30°C to obtain an inoculum ~ 2 × 108 CFU/ml.
2.4. Qualitative Estimation of Phosphate Solubilization Bacteria
Qualitative estimation of P solubilization was done by measuring the P solubilizing index (PSI). 10 μl of inoculum of each isolate was spotted on the NBRIP solid medium amended with 5.0 g/L TCP as a sole source of P and incubated at 30 °C for 14 days. The experiments were performed in four replicates and the sterile medium served as a control. PSI was calculated according to formula PSI = C+H/C, (C = Colony diameter; H = Halo zone diameter) [17].
2.5. Quantitative Estimation Of Phosphate Solubilization Bacteria
Quantitative estimation of P solubilization was determined by vanado-molybdat phosphoric method [18]. 50 ml of NBRIP broth medium adjusted to pH=7.0±0.2 before autoclaving was taken in 250 ml flask; then, it was inoculated with 200 μl of fresh inoculum and incubated in shaking condition at 120 rpm/min at 28°C for 7 days. Autoclaved and uninoculated NBRIP broth medium served as control. 2 ml of the cultures were taken and centrifuged at 10,000 rpm for 15 min. The content of soluble P was estimated by colorimetry; using the molybdenum blue method, optical density was measured on UV–VIS spectrophotometer at 882 nm. The pH of the NBRIP medium was also measured with a digital pH meter. All treatments were in triplicate.
2.6. Plant Growth-Promoting Traits of Phosphate Solubilization Bacteria
2.6.1. Siderophore Production
Siderophore production of each isolate was determined on Chrome-Azurol S (CAS) medium following the Universal Chemical Assay [19]. Bacterial strains (24 h fresh bacterial) were spotted on CAS plates and incubated at 28°C for 4 days. The production of siderophore was determined by the development of orange halo around the colonies. The experiments were performed in four replicates.
2.6.2. Indole-3-Acetic Acid Production
Based on their capacity to solubilize P, the selected PSBs were analyzed for IAA production. 200 μl of fresh bacterial cultures were inoculated in 30 mL of LB broth containing 0.1% L-tryptophan, and they were incubated in incubator Shaker at 28°C and 140 rpm/min for 72 h in dark. The bacterial cultures were centrifuged at 10,000 rpm for 10 min. After centrifugation, 2ml of supernatant was mixed with Salkowski reagent. After 30 min in dark, for pink color development, the optical density was measured at 530 nm using UV spectrophotometer. The amount of IAA produced was calculated by the standard graph of pure indole acetic acid [20].
2.6.3. Hydrogen Cyanide Production
Selected isolates were streaked on nutrient agar amended with 0.44% glycine. After that, a soaked filter paper in 2% Sodium carbonate and 0.5 ml picric acid solution was placed in the lid of the agar plate and sealed with parafilm. The plates were incubated for 4 days at 28°C. The turn of the yellow color to the brown one was considered as positive [21].
2.7. Identification of Isolated Strains
Nine isolates were selected based on their performance in TCP solubilization and their PGP trait. Crude DNA extractions from these selected isolates were prepared, and DNA Extraction was performed using the PureLink® Genomic DNA Mini Kit (Invitrogen, K1820-01) following the steps described by the manufacturer, adapted for Gram-negative bacteria. PCR amplification of the 16S rDNA of the bacterial strains was performed using the DreamTaq PCR Master Mix (Invitrogen), consisting of 22 mM Tris-HCl (pH 8.4), 55 mM KCl, 1.65 mM MgCl 2, 220 μM dGTP, 220 μM dATP, 220 μM dTTP, 220 μM dCTP, 22 U recombinant Taq DNA Polymerase / ml.
The universal primers 27F (sense) (5'AGAGTTTGAT CCTGGCTCAG-3') and 1492R (antisense) (5'-ACGGTTAC CTTGTTACGACTT-3') were used to amplify a 1500pb fragment, which corresponds to the genes of the bacterial 16S rDNA. PCR products were purified with the PureLinkTM Quick Gel Extraction & Purification combo kit (Invitrogen K220001), following the manufacturer's recommendations. Two-way sequencing was performed using primers 27F and 1492R. It was carried out according to the Sanger method adapted by the Big Dye Terminator V3 sequencing kit. Finally, it is the ABI3730 DNA sequences that allowed the automatic analysis of sequence reactions. The crude electropherograms were analyzed by MEGA7 software [22]. The consensus sequence from the sense and antisense raw sequences was obtained for each strain; then, it was compared to other sequences using the BLAST server (blast.ncbi.nlm.nih.gov) to determine their phylogenetic affiliation.
2.8. Statistical Analysis
Data were analyzed using SPSS 20 software, and the results were expressed as the means ± standard deviation of three replicates. Data were examined by ANOVA, and post-hoc mean comparison was performed by Duncan’s multiple range test at p ≤ 0.05.
| Location | Latitude | Longitude | Altitude (m) |
| L 1 | 32.75493458 | -6.85403284 | 683 |
| L 2 | 32.75526446 | -6.85307395 | 686 |
| L 3 | 32.75578241 | -6.85418146 | 679 |
| L 4 | 32.75619226 | -6.85484674 | 683 |
| L 5 | 32.75639581 | -6.85474465 | 684 |
| L 6 | 32.75636841 | -6.85437991 | 684 |
| L 7 | 32.75533956 | -6.85338013 | 689 |
| Samples | pH | Moisture (%) |
Organic Matter (%) |
Total Bacteria (UFC/mL) |
| S-BN | 7.84 ± 0.2 | 29.97 ± 1.02 | 7.84 ± 0.49 | 9.9 × 105 |
| S-BM | 7.1 ± 0.34 | 18.69 ± 0.9 | 7.1 ± 0.2 | 1.35 × 106 |
| S-BT | 7.84 ± 0.25 | 20.66 ± 0.90 | 2.97 ± 0.08 | 1.42 × 106 |
| S-ACY | 7.62 ± 0.11 | 10.13 ± 0.45 | 1.93 ± 0.09 | 2.06 × 106 |
| S-CB | 7.72 ± 0.09 | 13.80 ± 0.87 | 0.02 ± 0.004 | 1,48 × 105 |
| S-AC | 7.67 ± 0.5 | 10.04 ± 0.19 | 2.69 ± 0.08 | 4.87 × 106 |
| S-AT | 7.77 ± 0.67 | 6.08 ± 0.80 | 7.77 ± 0.23 | 1.75 × 106 |
3. RESULTS
3.1. Selection and Purification of Phosphate Solubilization Bacteria
Data on the physicochemical analysis of the samples of the PSS are represented in Table 2. The distribution of total PSB isolated from seven phosphate solid sludge samples is shown in Table 3. PSBs isolation and selection were achieved in NBRIP solid medium. The strains were selected according to morphological difference colonies and TCP solubilization halos. Although the halo formation around the colonies provided the first qualitative indication of PSB, the colonies which were developed without forming a halo were also isolated to test their performance in the solubilization. As a first step, 150 isolates were selected; these isolates were transplanted on NBRIP medium 3 times. The isolates that possessed the ability to form a halo were selected. Out of 150 isolates, 27 strains were selected from the different PSS samples as follows: where 29.63% were isolated from S-BM, 22.22% from S-CB, 22.22% from S-AT, while 18.51%, 3.7%, 3.7% were obtained from S-BN, S-AC, S-ACY respectively, and no PSB isolates were selected from S-AT sample (Table 3).
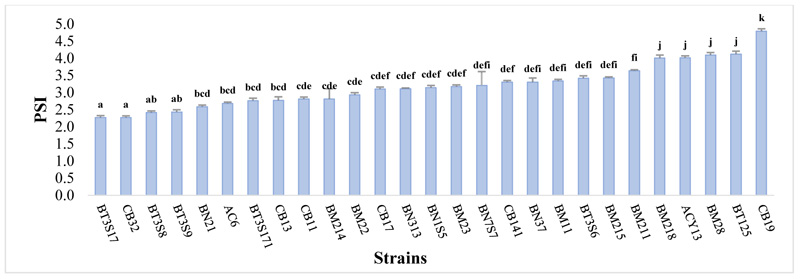
3.2. Qualitative Estimation of Phosphate Solubilization Bacteria
A significant difference was observed in all strains tested (P ≤ 0.05) Fig. (1). The soluble phosphate in NBRIP sold medium ranged from 2.27≤ SPI ≤4.79, and the appearance of the halo zone in all strains was remarked after the 3rd day of incubation. Maximum PSI was noticed in CB19 with 4.79±0.57, followed by BT125 (4.12±0.49), BM28 (4.11±0.25) and ACY13 (4.02±0.12). Moderate solubilization index was observed in BT3S17 (2.27±0.24) and CB32 (2.27±0.14).
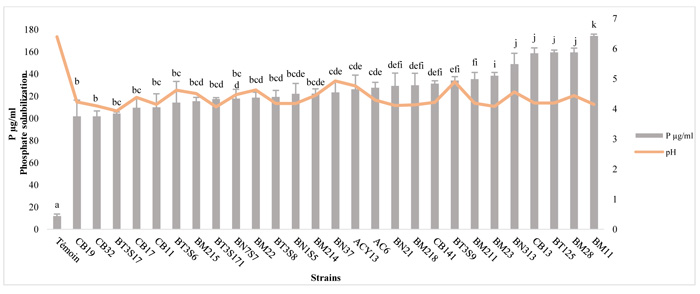
3.3. Quantitative Estimation of Phosphate Solubilization Bacteria
The amounts of Soluble-P and pH variation are presented in Fig. (2). The TCP solubilization in NBRIP liquid medium, by the strains tested, was accompanied by a significant decrease in pH. The soluble-P concentration in the medium ranged between 101.91 μg/ml and 174.33 μg/ml with significant variations among different isolates (p ≤ 0.05). Maximum TCP solubilization in NBRIP liquid medium was observed in BM11 (174.33±12.5μg/ml), followed by BT125 (159.48±1.8 μg/ml), BM 28 (159.48±1.7 μg/ml) and CB13 (158.73±3.6 μg/ml), and Modest P solubilization was observed in CB32 (101.91±9.6 μg/ml). In the control, 12±1.6 μg/ml of P was solubilized in the culture medium.
3.4. Plant Growth-Promoting Traits of Phosphate Solubilization Bacteria
3.4.1. Siderophore production
Siderophore production was carried out on solid CAS blue agar; the development of orange halo around the colonies represents iron chelation by the isolate in the medium. The totality of the tested isolates confirmed the secretion of siderophore (Table 4). 48.14% of the isolates showed a high production of siderophore, while 29% of the isolates showed a medium performance, and 22.22% showed a poor efficiency.
3.4.2. IAA Production
All selected isolates demonstrated an important production of IAA with a significant difference between strains (p ≤ 0.05) (Table 4). Highest IAA production was reported in BT3S171 with 43.80±0.39 μg/ml, followed by BM218 (20.53±0.03 μg/ml) and BM215 (15.96±0.20 μg/ml), and the lowest production of IAA was detected in BN313 with 1.37±0.09 μg/ml.
| Samples | Isolates | Strains % |
| S-BN | BN313, BN1S5, BN7S7, BN21, BN37 | 18.51 |
| S-BM | BM11, BM28, BM22, BM23, BM21, BM214, BM215, BM218 | 29.63 |
| S-BT | BT125, BT3S17, BT3S171, BT3S6, BT3S8, BT3S9 | 22.22 |
| S-ACY | ACY13 | 3.7 |
| S-CB | CB17, CB11, CB19, CB141, CB13, CB32 | 22.22 |
| S-AC | AC6 | 3.7 |
| S-AT | - | 0 |
| Strains | IAA (µg/mL) | Siderophore | Hydrogen Cyanide |
| Control | 0.12 ± 0.01a | - | - |
| BT125 | 3.90 ± 0.04 igk | +++ | ++ |
| BM218 | 20.53 ± 0.03r | + | - |
| BM28 | 7.69 ± 0.03 n | ++ | - |
| BM211 | 2.66 ± 0.03 bcdefig | + | - |
| BN37 | 11.47 ± 0.01p | ++ | - |
| BM11 | 2.05 ± 0.08 bcd | ++ | - |
| BN313 | 1.37 ± 0.09 b | +++ | - |
| CB13 | 8.73 ± 0.19 n | ++ | - |
| CB141 | 2.53 ± 0.04 bcdefi | +++ | - |
| BN21 | 3.40 ± 0.04 fig | +++ | - |
| BT3S6 | 1.64 ± 0.05 bc | ++ | - |
| BM22 | 11.49 ± 0.61o | + | - |
| AC6 | 4.77 ± 0.16 lm | +++ | + |
| ACY13 | 3.12 ± 0.13 cdefig | ++ | - |
| BM23 | 4.06 ± 0.04 gkl | + | - |
| BN1S5 | 5.71 ± 0.05 m | + | +++ |
| CB17 | 2.86 ± 0.33 fig | +++ | +++ |
| BT3S9 | 2.76 ± 0.03 cdefig | +++ | - |
| BT3S8 | 3.22 ± 0.05 defig | +++ | ++ |
| BN7S7 | 8.19 ± 0.15 n | + | +++ |
| CB19 | 3.17 ± 0.03 efig | ++ | - |
| BT3S17 | 1.90 ± 0.03 bcde | +++ | - |
| BT3S171 | 43.80 ± 0.39 s | +++ | - |
| CB32 | 3.12 ± 0.05 efig | +++ | - |
| CB11 | 4.99 ± 0.13 klm | +++ | - |
| BM214 | 2.38 ± 0.01bcdef | ++ | - |
| BM215 | 15.96 ± 0.20 q | +++ | - |
| Strain | Gram | 16S rRNA Gene Sequence Lengh | Accession N° | Species | % Gene Identity |
| BM11 | Gram | 1350bp | NR_024644.1 | Serratia rubidaea strain JCM1240 | 99.70% |
| BM28 | Gram | 1350bp | NR_148649.1 | Enterobacter bugandensis strain 247BMC | 99.33% |
| BM215 | Gram | 1300bp | NR_148649.1 | Enterobacter bugandensis strain 247BMC | 99.23% |
| CB19 | Gram | 1350bp | NR_114505.1 | Pantoea agglomerans strain ATCC 27155 | 98.00% |
| BN313 | Gram | 1350bp | NR_156986.1 | Pseudomonas lactis strain DSM 29167 | 99.85% |
| BT125 | Gram | 1350bp | NR_116299.1 | Pseudomonas brassicacearum subsp. Neoaurantiaca strain CIP 109457 | 99.78% |
| BT3S171 | Gram | 1300bp | NR_104928.1 | Pantoea stewartii subsp. Indologenes strain CIP 104006 | 98.59% |
| CB13 | Gram | 1300bp | NR_114505.1 | Pantoea agglomerans strain ATCC 27155 | 98.00% |
| BM218 | Gram | 1350bp | NR_114505.1 | Pantoea agglomerans strain ATCC 27155 | 98.00% |

3.4.3. Hydrogen Cyanide Production
Out of the 27 isolates, only six isolates (22.2%) showed HCN production after 4 days of incubation (Table 4). Maximum HCN production was observed in BN1S5, BN7S7 and CB17, followed by BT125 and BT3S8. AC6 was the least efficient isolate in HCN production.
3.5. Identification and Characterization of the Isolated Strains
The identifications of PSBs strains, based on 16S rRNA sequences, are presented in Table 5. Among 27 bacterial strains, nine strains (BM11, BM28, BM215, BM218, CB13, C19, BN313, BT125, BT3S171) were selected for molecular characterization, based mainly on their solubilization capacity and also on their PGP traits: IAA secretion, siderophore production and the HCN released from genetic analyses; we identified four genera: Pseudomonas, Serratia, Pantoea and Enterobacter. Based on the analysis of the 16S rDNA partial sequence, the strain BM11 was identified as Serratia rubidaea. Strains BM28 and BM215 were identified as Enterobacter bugandensis. Moreover, three strains were identified as Pantoea agglomerans. Strains CB19, CB13, and BM218, BT3S171 were identified as Pantoea stewartii subsp. Indologenes. BT125 was identified as Pseudomonas brassicacearum supsp.neoaurantiaca, and strain BN313 was identified as Pseudomonas lactis (Fig. 3).
4. DISCUSSION
The objective of our study was to isolate PSB from phosphate solid sludge samples, based on the NBRIP medium with TCP as the sole source of phosphorus. From 150 isolates, we selected 27 strains qualified as phosphate solubilizers, after evaluating their ability to solubilize TCP and their PGP traits. We chose 9 isolates for molecular characterization.
It has been found that P is mandatory for plant growth, and its deficiency limits plant development. Even though chemical fertilizers are added to the soils, plants can only use low phosphatic fertilizer quantity because of the immobilization of P. Thus, the selection of highly efficient PSB is very important; it will practically raise P in plant rhizosphere. Various PSB have been isolated from different soil and rhizosphere [23]. As a consequence, PSB can be considered as a plant growth-promoting rhizobacteria, which are commonly regarded as biofertilizers [24, 25]. Also, the presence of PSB in soils will be judged as a positive indicator of using the bacteria as biofertilizers for crop production [23, 26]. In our study, the selected isolates showed high efficiency of solubilization, the soluble-P concentration in the NBRIP medium ranged between 101.91 μg/ml and 174.33 μg/ml. This result is in accordance with previous studies, the solubilization of TCP in their experiments was from 41.20 to 119.95 μg/ml in the NBRIP medium and from 30 to 246 μg/ml respectively [27, 28]. Phosphate solubilization ability has been considered as a principal criterion for isolating highly efficient PSB strains from phosphate solid sludge, but we have also focused on other PGP criteria. Numerous studies have been conducted to isolate effective PGPR according to various criteria, such as IAA and siderophore production or HCN production [29-31]. IAA, siderophore and HCN production are an indicator of PGP Rhizobacteria [29, 32]. IAA production by bacteria isolated from the soil has previously been reported [33-35] as an important phytohormone, and as a regulatory molecule in plant development [32]. In the present study, all the isolates produced IAA within the range of 43.80 μg/ml – 1.37 μg/ml (p ≤ 0.05), which indicated a significant variability among isolates for IAA production. Moreover, another important trait of PGPR is the production of siderophores which influences plant growth. They are produced by numerous types of bacteria in response to iron deficiency, and they play a vital role in determining the competitive fitness of bacteria to colonize plant roots and to compete for iron with other microorganisms in the rhizosphere [36]. In our study, all strains produced siderophore with a significant difference between the strains; HCN production by rhizobacteria has been postulated to play an important role in the biological control of pathogens. HCN is volatile, a secondary metabolite that inhibits the growth of pathogen due to inhibitor of metal enzymes, particularly cytochrome C oxidases in an electron transport system [37]. In this research, it has been found that from 27 isolates, only 22.2% produced HCN. These results closely correlate with the study [27] which reports that 83.33% of the strains produce IAA, while 66.66% of strains have HCN and siderophore producers. The [38] study also reveals that 95% of their isolated PSB strains are IAA producers, 100% of strains produce siderophores and 67.5% of strains are HCN producers. In the literature, numerous bacterial strains are considered as PSBs, and among them, those from genera are Pseudomonas, Serratia, Pantoea, Alcaligenes, Enterobacter, Acinetobacter, Arthrobacter, Azospirillum, Bacillus, Burkholderia, Erwinia, Flavobacterium and Paenibacillus. Our selected strains belong to four genera which have been deemed as PSB: Pseudomonas, Serratia, Pantoea and Enterobacter. These results are congruent with those of [10, 39-42].
The use of PSB as biofertilizers would strengthen the use of chemical fertilizers and promote plant growth. Taken together, the selection of an efficient PSB strain as inoculants and qualifying them as biofertilizers should be based not only on laboratory experiments and greenhouse trials but also on field experiments. Further studies should focus on practical applications of PSB in the field.
CONCLUSION
A total of 27 strains were isolated in this study, among which 9 were identified; they belong to the genus: Pseudomonas, Serratia, Pantoea, and Enterobacter. Results showed that P solubilization by PSB in the NBRIP media performed differently and most of the strains displayed plant growth-promoting traits, thereby indicating that they have a role in enhancing plant growth. Further studies should be carried out, namely the production of organic acids by those strains. We can also test them on crops as biofertilizers in greenhouse.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
This work is conducted under a project sponsored by OCP Foundation under the number BIO-ELG-01/2017.
CONFLICT OF INTEREST
The author declares no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors would like to acknowledge the support through the R&D Initiative – Appel à projets autour des phosphates APPHOS.


